 Most linear-time string searching algorithms are tricky to implement, and require heavy preprocessing of the pattern before running the search. This article presents the Rabin-Karp algorithm, a simple probabilistic string searching algorithm based on hashing and polynomial equality testing, along with a Python implementation. A streaming variant of the algorithm and a generalization to searching for multiple patterns in one pass over the input are also described, and performance aspects are discussed.
Most linear-time string searching algorithms are tricky to implement, and require heavy preprocessing of the pattern before running the search. This article presents the Rabin-Karp algorithm, a simple probabilistic string searching algorithm based on hashing and polynomial equality testing, along with a Python implementation. A streaming variant of the algorithm and a generalization to searching for multiple patterns in one pass over the input are also described, and performance aspects are discussed.
The algorithm is probabilistic in that it doesn’t always return correct results; more precisely, it returns all valid matches and (with reasonably small probability) a few incorrect matches (algorithms such as this one that tend to be over-optimistic in reporting their results are usually said to be true-biased).
Algorithm
Given a string s of length \(n\), and a pattern of length \(k\), the algorithm computes rolling checksums for both the pattern (“needle”) and consecutive substrings of length \(k\) of the searched string (“haystack”). Every time a substring’s checksum equals that of the pattern, a match is reported.
matches = {}
pattern_checksum = checksum(pattern)
substring_checksum = checksum(s[0 : k[)
for position from 0 to n - k - 1 do
update substring_checksum to checksum(s[position : position + k[)
if substring_checksum = pattern_checksum then
add position to matches
return matches
Checksum functions
Formally, checksum functions map fixed-length strings to numbers, with good checksum functions satisfying two properties: first, collisions (two different strings being mapped to the same checksum) must be rare — indeed, the higher the collision rate, the more likely incorrect matches are to turn up. Second, checksums must be computed in a way that allows for efficient updating: that is, the value of the checksum of s[i+1:i+k+1[ should be quickly derivable from that of s[i:i+k[.
Polynomial hash functions
One easily implemented such hash function is based on polynomials. Given a list of \(k\) numbers \(a_0, \ldots, a_{k-1}\), we define a polynomial ((Using \(\sum_{j < k} a_j X^{(k-1)-j}\) instead of \(\sum_{j < k} a_j X^j\) yields a cleaner rolling checksum update formula in the next section.))
$$ H(a)(X) = a_0 X^{k-1} + a_1 X^{k-2} + \ldots + a_{k-2} X + a_{k-1} = \sum_{j < k} a_j X^{(k-1)-j} $$
Assuming that each character of a string \(s\) can be matched to numbers, we then define the checksum of a substring s[i:i+k[ of length \(k\) of \(s\) as \(H(s[i:i+k[)(X) = \sum_{j < k} s_{i + j} X^{(k-1)-j}\). ((The weakness of this approach is that it may require a normalization pass to successfully process some strings using extended character sets; indeed, some Unicode characters can be represented in multiple ways: the capital A with diaeresis character Ä, for example, can be represented either as U+00C4 (Ä), or U+0041 (A) + U+0308 (¨) yielding Ä. Similarly, Unicode includes support for ligatures — that is one can encode the pair “fi” as either U+0066 U+0069 “fi” or the ligature U+FB01 “fi”. MSDN and Wikipedia have more details on this.))
Polynomial identity testing
As it stands, the mapping from s[i:i+k[ to \(H(s_{[i:i+k[})\) is bijective, and the problem of checking whether s[i:i+k[ matches a pattern p can thus be reformulated as checking whether \(H(s_{[i:i+k[})(X) = H(p)(X)\). This problem is usually called polynomial identity testing, and can be deterministically solved in time \(O(k)\) by comparing polynomials coefficient-wise.
Allowing for infrequent errors, however, takes the complexity of this process down to probabilistic \(O(1)\): instead of comparing two polynomials \(P\) and \(Q\), one can pick a random value \(a\) and compare the values of \(P(a)\) and \(Q(a)\), accepting if and only if \(P(a) = Q(a)\) (since values of \(P(a)\) and \(Q(a)\) may be too large to fit in standard-sized integers, computations are generally made modulo a certain large prime modulus \(m\)). The reasoning is that if indeed \(P=Q\) then the test will always accept \(P\) and \(Q\) as equal, while if \(P \not= Q\) then the probability of hitting a bad witness and incorrectly accepting \(P\) and \(Q\) as equal is low.
More precisely, if \(P\) and \(Q\) are polynomials of degree \(≤ k\), and \(m\) is a prime number larger than the largest coefficient of \(P – Q\), then the probability of hitting a root of \(P-Q\) is lower than \(k/m\) if \(P\) and \(Q\) differ ((Given a prime number \(m\), \(\mathbb{Z}/(m\mathbb{Z})\) is a finite field and a \(k\)-polynomial whose coefficients fall in the range \(0 \ldots m-1\) has at most \(k\) roots in \(\mathbb{Z}/(m\mathbb{Z})\).)).
Picking a large enough random prime modulus \(m\) and a random seed \(a \in \{ 0 \ldots m – 1 \} \) therefore allows for the safe definition of $$h_i = H(s_{[i:i+k[})(a) \mod m = \sum_{0 ≤ j < k} s_{i + j} a^{(k-1)-j} \mod m$$ as the checksum of s[i:i+k[.
Rolling checksum update
This definition allows for the checksum \(h_{i+1}\) of s[i+1:i+k+1[ to be derived from that of s[i:i+k[ (\(h_i\)) in constant time (provided the value of \(a^k\) is pre-computed once and for all before running the algorithm), using the formula \(h_{i+1} = a · h_i – s_i a^k + s_{i+k}\). Indeed,
$$
\begin{align}
h_i = H(s_{[i+1:i+k+1[})(a) &= \sum_{0 ≤ j < k} s_{i + 1 + j} a^{(k-1)-j}\\
&= \sum_{1 ≤ j < k + 1} s_{i + j} a^{(k-1)-(j-1)}\\
&= \sum_{0 ≤ j < k} s_{i + j} a^{(k-1)-j+1} – s_i a^k + s_{i+k}\\
&= a · H(s_{[i:i+k[})(a) – s_i a^k + s_{i+k}\\
&= a · h_i – s_i a^k + s_{i+k}
\end{align}
$$
Python implementation
import random # See the implementation notes below for details. "(→ n)" indicates a footnote. def checksum(string, seed, modulus, length): """Computes the checksum of string[0 : length+1]""" checksum = 0 for pos in range(0, length): checksum = (checksum * seed + ord(string[pos])) % modulus return checksum def string_search(haystack, needle): """Yields the list of positions p such that haystack[p : p + len(needle) - 1] == needle""" LARGEST_CODEPOINT = 0x10FFFF # (→ 1) # Sanity check if len(haystack) < len(needle): return # Initialization needle_length = len(needle) haystack_length = len(haystack) modulus = 0xB504F32D # (→ 2) seed = random.randint(0, modulus - 1) seed_k = pow(seed, needle_length, modulus) # Pre-compute a^k needle_checksum = checksum(needle, seed, modulus, needle_length) substr_checksum = checksum(haystack, seed, modulus, needle_length) # (→ 3) # String processing for pos in range(0, haystack_length - needle_length + 1): if needle_checksum == substr_checksum: # (→ 4) yield pos if pos < haystack_length - needle_length: # (→ 5) substr_checksum = (seed * substr_checksum - ord(haystack[pos]) * seed_k + ord(haystack[pos + needle_length])) % modulus
Implementation notes
- LARGEST_CODEPOINT = 0x10FFFF
- Characters are mapped to polynomial coefficients using the python
ordfunction, which returns the Unicode code point of any character. The largest such code point is 0x10FFFF. - modulus = 0xB504F32D
-
Checksum polynomials coefficients are smaller than
LARGEST_CODEPOINT, and as the modulus must exceed all checksum polynomial coefficients it must exceedLARGEST_CODEPOINT((Using a modulus smaller thanLARGEST_CODEPOINT+1would introduce spurious polynomial equalities; for example, \(X^3 + 3X^2 + 1 \not= X^3 + 1 \mod 7\), but \(X^3 + 3X^2 + 1 = X^3 + 1 \mod 3\).)). Further requiring that the modulus be as large as possible helps reduce the probability of checksum collisions, and hence of spurious matches. Finally, enforcing that it be prime bounds the number of integer roots of degree-k polynomials.0xB504F32Dwas chosen as it is the largest value satisfying these requirements whose square fits in a 64-bit signed integer. ((This definition definition could be rewritten as follows, assuming the availability of a the availability of arandom_prime(n)function to sample a random integer from the range \(n, \ldots, 2n – 1\):modulus = random_prime( max(2**30.5, LARGEST_CODEPOINT + 1) ))) - substr_checksum = checksum(haystack, seed, modulus, needle_length)
- Before starting the rolling computation,
substr_checksumis initialized to the checksum of \(s[0:k[\). - if needle_checksum == substr_checksum:
- To prevent checksum collisions from yielding incorrect matches, one could explicitly verify matches ((If such a verification is implemented, then this algorithm is exactly the Rabin-Karp string searching algorithm)) by defining
def verify(haystack, pos, needle): return all(haystack[pos+k] == needle[k] for k in range(0, len(needle)))
and replacing the checksum test with
if needle_checksum == substr_checksum and verify(haystack, needle, pos):
This slows down the algorithm a bit, however, taking its worst-case performance to \(O(nk)\) ((This worst-case bound is attained when both the haystack and the needle are composed of only one character, e.g. when searching for \(aaa\) in \(aaa \ldots aaa\))) — in general, however, the performance remains good as long as the pattern is not redundant. ((One could furthermore argue that the cost of this additional post-checking step is not significant, as string searching programs like
grepgenerally display matches anyway, taking linear time in the length of the pattern for each match. Since verifications only occur on matches detected by checksum comparison, all that remains to be checked is that spurious matches are rare enough to not adversely affect execution time.)) - if pos < haystack_length – needle_length:
- This check prevents an out-of-bounds access after the last comparison.
- 32- vs 64-bits performance
- The sample code above manipulates relatively large numbers (close to \(2^{31.5}\)), with intermediate calculation results reaching close to \(2^{63}\). Such calculations are fast on 64 bit machines, but will trigger promotion to Python’s arbitrary-size integers on 32-bit machines, thus causing the code to perform more poorly. Redefining
modulusas0xB501would do, at the cost of increasing the rate of erroneous matches.
Incorrect matches: theoretical bounds and experimental results.
As previously noted, the probability of a spurious match turning up at position \(i \in \{0, \ldots, n-1\}\) is lower than \(k/m\) (\(n\) denotes the length of the haystack, \(k\) that of the needle, and \(m\) is the modulus). The probability of at least one incorrect match turning up at any position in the string, however, is higher — indeed, in the general case, it can only be shown to be lower than \(n\) times probability of an error popping up at an arbitrary given position ; that is, \(n · k/m\). An ideal implementation would choose \(m\) to be as large as \(n · k\) times a certain large constant \(C\), thus taking the probability of one or more incorrect matches popping up down to \(1/C\).
Real-life implementations, however, are constrained by the size of machine integers (avoiding automatic promotion to Python’s arbitrary-sized integers is crucial to ensuring acceptable performance). The implementation above therefore chooses the largest safe modulus value on a 64-bits architecture, 0xB504F32D ((Python standard integers can go up to \(2^{63} – 1\) before being promoted to arbitrary-precision integers)). This takes the probability of obtaining one or more incorrect matches down to about \( n · k / 2^{32} \), which is below \(0.5\%\) for typical string searching problems ((For example, searching for a 10-character string like “Petersburg” in the plain text version of Dostoyevsky’s Crime and punishment (~ 1.1 million characters long).)).
The culprit here is the checksum update calculation, and in particular the seed * substr_checksum multiplication, which yields values as big as the square of the modulus. Unfortunately there is no way (that I am aware of) to perform the multiplication and integer division that does not resort to large integers if the intermediary multiplication result exceeds the size of machine integers. And it gets even worse on 32-bit machines, where the modulus must be chosen smaller than \(2^{15} – 1\).
Low-level optimizations, however, allow for the use of larger moduli:
- In C, one bit can be saved by using unsigned integers, pushing the modulus up to
0xFFFFFFFB. In that case, special care must be taken in the checksum update code, so that the subtraction does not cause wrapping. The naïve implementation, shown below, performs incorrectly:substr_checksum = (seed * substr_checksum - outgoing * seed_k + incoming) % modulus
Specifically, if the result of the
seed * substr_checksum - outgoing * seed_kis below 0, then the value is wrapped modulo(1 << 64) - 1((Wrapping behaviour is defined in sub-clause 6.2.5 paragraph 9 of the C Standard: “A computation involving unsigned operands can never overflow, because a result that cannot be represented by the resulting unsigned integer type is reduced modulo the number that is one greater than the largest value that can be represented by the resulting type”.)). Instead, the correct calculation is as show below:uint64_t mul = (seed * substr_checksum) % modulus; uint64_t sub = ( (modulus - outgoing) * seed_k ) % modulus; substr_checksum = (mul + sub + incoming) % modulus;
- In 64-bits assembler, using the unsigned multiplication
MULinstruction to calculate the product above yields a 128-bits result stored in theEDX:EAXregisters, whose remainder in the division by a 64-bits modulus can be calculated using theDIVinstruction (MODon HLA).
More importantly, this \(n · k/m\) bound on the error rate is only theoretical, and the effective error rate on real-life texts will often be much lower. Experimental results are hard to obtain given the time it takes to search through large files, but searching 200 times for a 5000-characters random string in Project Gutenberg’s pg2554 (Crime and punishment), didn’t return a single false positive, though the theoretical bound that can be derived from the formula above regarding the probability of getting at least one erroneous match in such an experiment was above one.
In a second experiment, I searched for the string TATAAA (TATA box consensus sequence, length 6) in the genome of a female platypus (length 1 684 663 807). Again, no incorrect matches were found.
Streaming
One nice property of the algorithm above is that it does not require that the entire string be loaded into memory; instead, it just requires a buffer whose length equals that of the pattern being searches for. This makes it especially suitable for looking for relatively small patterns in huge strings, like gene patterns in DNA strings. In this particular case verifying each match is especially advisable, given that the input stream is discarded after being processed, making it impossible to verify matches after the algorithm has finished running. Plus, the enormous length of the input does increase the probability of an incorrect match popping up at some point.
The modifications required to allow for streaming are presented below:
import random, itertools def verify(match, needle): return all(match[k] == needle[k] for k in range(0, len(needle))) def checksum(input, seed, modulus): checksum = 0 for byte in input: # (→ 1) checksum = (checksum * seed + byte) % modulus return checksum def stream_search(stream, needle): """Yields the list of positions p such that stream[p : p + len(needle) - 1] == needle""" # Initialization needle_length = len(needle) modulus = 0xB504F32D seed = random.randint(0, modulus - 1) seed_k = pow(seed, needle_length, modulus) # Pre-compute a^k prefix = list(itertools.islice(stream, needle_length - 1)) # (→ 2) needle_checksum = checksum(needle, seed, modulus, needle_length) substr_checksum = checksum(prefix, seed, modulus, needle_length - 1) # Stream processing pos = 0 substr = collections.deque(prefix) # (→ 3) for incoming in stream: substr.append(added) outgoing = substr.popleft() if pos > 0 else 0 # (→ 4) substr_checksum = (seed * substr_checksum - outgoing * seed_k + incoming) % modulus if needle_checksum == substr_checksum and verify(substr, needle): yield pos pos += 1
Streaming implementation notes
- for byte in input:
- We now process
bytesobjects instead ofstringobjects, as we are mostly going to process huge byte streams loaded from files or from a network. - prefix = list(itertools.islice(stream, needle_length – 1))
- The
itertools.islice(stream, n)function consumesnvalues from a stream. We only take theneedle_length - 1first ones here, and the first round of update will complete the checksum calculation. - substr = collections.deque(prefix)
- Values corresponding to the substring whose checksum is being calculated are stored in a sliding window implemented as a double-ended queue. This window is required as the first and last values that it contains are needed to update the rolling checksum.
- outgoing = substr.popleft() if pos > 0 else 0
- This conditional check prevents
popleftfrom being called on an incomplete window in the first rolling checksum update step. This is needed because on the first iteration of the loop the checksum is incomplete, and the sliding window is not full yet.
Performance
As it stands, the algorithm presented above performs rather poorly for two reasons:
- Implementation weaknesses: Using iterators to loop over the input string introduces a heavy overhead in Python
- Algorithm weaknesses: Extracting a modulus is a slow operation. Plus, the algorithm used processes all characters; CPython’s default implementation, in contrast, uses a deterministic algorithm based on two popular string searching algorithms, Boyer-Moore and Horsepool, that skips parts of the input.
Though it performs acceptably, the python implementation of this algorithm is therefore no match for the highly optimized built-in string.find CPython function. The first culprit pointed out above can easily be fixed by moving to another, more low-level language ((Indeed, a C implementation of this simple algorithm runs only about 20 times slower than Python’s heavily-optimized routines)); the second, which is directly related to the algorithm being used, is much harder to mitigate.
Using a non-prime modulus is sometimes advocated; in that case, the modulus most often chosen is generally \(2^{64}\) (or whatever the size of machine integers is (+1), the idea being that C wrapping semantics guarantee that the modulus operation will be performed almost for free by the hardware; in that case the modulus operations can be removed altogether, but care must be taken to ensure a proper choice of seed, so as to minimize the probability of checksum collisions. This yields an experimental speedup of about 4 times in C; the resulting code is in the order of 5 to 10 times slower than the optimized CPython routine, but it is deterministic, and hence vulnerable to collision-based attacks.
Multi-pattern search
Where Rabin-Karp does shine is when searching for multiple, same-length patterns. With trivial modifications, the algorithm above can indeed be extended to match the current substring’s checksum against a list of pre-computed pattern checksums. Provided this lookup operation is implemented efficiently enough, Rabin-Karp becomes an efficient alternative to repeatedly running a more efficient single-pattern search algorithm.
The modified pseudo-code is presented below:
matches = {}
patterns_checksums = [checksum(pattern) for pattern in patterns]
substring_checksum = checksum(s[0 : k[)
for position from 0 to n - k - 1 do
update substring_checksum
if substring_checksum belongs to pattern_checksum then # (→ 1)
add position to matches
return matches
All that remains is to find a proper implementation of the belongs to (→ 1) operation. Hash-tables, though they might seem appropriate, require a large amount of memory to not introduce extra collisions ((The streaming implementation presented above only requires keeping the last \(k\) tokens in memory, and is hence rather memory-efficient)), and they introduce a slight performance overhead. Instead, sorting patterns checksums and binary-searching the resulting array does not require any additional memory, and introduces very low overhead for reasonable patterns count.
The following graph demonstrates the effects of using this modified algorithm to search for large numbers of patterns. Four implementations are compared:
- Naive python, C implementations:
for pattern in patterns: if haystack.find(pattern) > 0: return True
for (int pattern_id = 0; pattern_id < patterns_count; pattern++) if (strstr(haystack, patterns[pattern_id])) return true;
Execution time grows linearly with the number of patterns. Useless for more than 50 patterns. Requires multiple passes on the input.
- Regular-expressions–based python implementation:
regexp = "|".join(patterns) if re.search(regexp, haystack): return True
Execution time also grows linearly with the number of patterns, but the growth is much slower, making this solution practical for patterns counts under 5000. ((Ideally, though, the regular expression being generated would be transformed into a deterministic automaton whose performance would only marginally depend on the number of patterns (incidentally, this is the basis of the Aho-Corasick algorithm). Unfortunately, Python doesn’t do that by default.)) Requires only one pass on the data. Requires sanitization of the input patterns for the regular expression to compile properly.
- C++ multi-patterns Rabin-Karp implementations:
Based on the description above. Execution time grows logarithmically in the number of patterns. Much slower than other techniques for small patterns count, but reasonable for patterns count above 50, and optimal above ~2000/5000 depending on implementation-dependent details. Requires only one pass. Using a hardware-based modulus (wrapping at \(2^{64}\)) does yield a significant performance improvement.
The following graph shows the respective performance of these implementations.
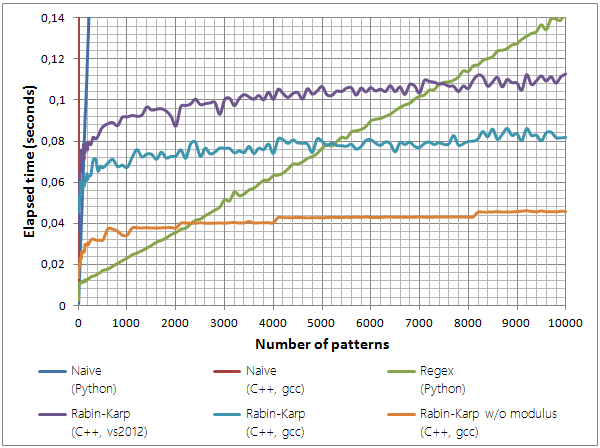
Multi-pattern search performance. While naive implementations quickly become impractical, the regex-based implementation yields pretty good performance. It is eventually superseded, however, by Rabin-Karp–based implementations, notably by the hardware modulus implementation. (Experimental settings: searching for random strings of length 11 in pg2554.)
Conclusion & further reading
Although its performance if only middling on single-pattern string searching problems, the randomized algorithm presented above shines by its conceptual simplicity, its ease of implementation, and its performance on multi-string searching problems. One of its most successful applications is the implementation of similarity-detection tools, which search for a large number of known sentences in newly submitted texts (articles, exams, …).
The original paper by Rabin & Karp (1986) is a fascinating read; it includes much more details on the algorithm’s expected performance in various settings, and generalizations to a broader class of problems. Further results on the average-case complexity of this algorithm were presented by Gonnet & Baeza-Yates in 1990.
Another approach to the multi-string matching problem, that relies on the remarks made above on converting regular expressions to finite automaton, was presented by Aho & Corasick in 1975. It basically preprocesses the patterns input set to construct a finite deterministic automaton, thus achieving one-pass, linear-time matching of all patterns in the set. Its implementation, however, is much trickier than that of Rabin-Karp.
Have you ever had to use string searching functions other than that provided by the standard library of you programming language? Which algorithms did you use?